 | Research |

''
''
Create New World of Bioactive Synthetic Molecules
New ways to use, New Shapes, New sizes. †
As biological processes all stem from chemical events, it should be possible to understand or manipulate biological events by using chemistry. As chemical biologists our challenge is to discover or design unique organic molecules, 'super tools' that modulate fundamental processes in human cells. Using these 21st century super tools we can explore complex cellular events. Our job is creation: designing a new world of bioactive synthetic molecules, with unique and novel uses, forms and sizes. The future for small-molecule applications is open before us, in a range of fields, including future drug discovery and cell therapy. Our 21st century technology and skills will revolutionize the fight against the most difficult diseases and medical challenges of the age.
ケミカルバイオロジー~化学を起点とした生物学 †
ケミカルバイオロジーとは、化学を起点とした生物学です。生命の営みはせんじつめれば化学反応でできています。逆に化学を使って生命現象を理解したり、操ることができるはず。私たちの研究室では、生き物やヒト細胞にユニークな効果を及ぼす独自な有機化合物を見つける、もしくはデザインし、それらを道具として生命現象を探究・操作してきました。生物の仕組みは複雑ですが、有機化合物を起爆剤として用いることで、新たな切り口で生物を研究したり操作することができます。私たちの大きな研究目標は、生理活性合成化合物の新しい世界を切り開くことです。生理活性化合物の新しい利用法、新しいカタチをした生理活性化合物、新しいサイズの生理活性化合物など、アイデアを与えることができればと思うのです。こういったアイデアは、未来の創薬の考え方や化合物の細胞治療への利用などに結びつくはずです。
''
''
Selected Publications †
Covalent Plant Natural Product that Potentiates Antitumor Immunity †
Takemoto, M., Delghandi, S., Abo, M., Yurimoto, K., Odagi, M., Singh, V., Wang, J., Nakagawa, R., Sato, S., Takemoto, Y., Farrag, A., Kawaguchi, Y., Nagasawa, K., Honjo, T., Chamoto, K., Uesugi, M. †
J. Am. Chem. Soc. 147(3), 2902–2912 (2025)
https://doi.org/10.1021/jacs.4c17837

Chemoproteomic Identification of Spermidine-Binding Proteins and Antitumor-Immunity Activators †
Vaibhav Pal Singh, Shuhei Hirose, Misao Takemoto, Asmaa M. A. S. Farrag, Shin-ichi Sato, Tasuku Honjo, Kenji Chamoto*, and Motonari Uesugi †
J. Am. Chem. Soc. 146(24), 16412–16418(2024)
https://doi.org/10.1021/jacs.3c14615

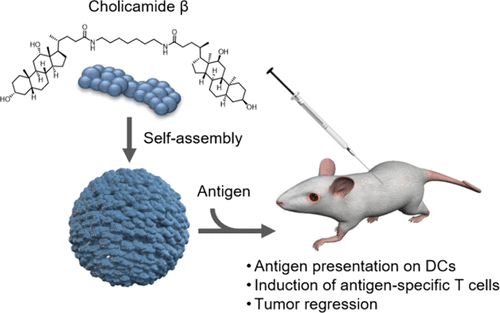
Identification of a Self-Assembling Small-Molecule Cancer Vaccine Adjuvant with an Improved Toxicity Profile †
Zhuo, S., Noda, N., Hioki, K., Jin, S., Hayashi, T., Hiraga, K., Momose, H., Li, W., Zhao, L., Mizukami, T., Ishii, K., Li, Y., Uesugi, M. †
J. Med. Chem. 66(18), 13266–13279 (2023)
https://doi.org/10.1021/acs.jmedchem.3c01252
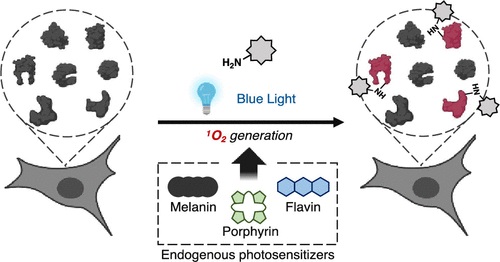
Chemoproteomic Identification of Blue-Light-Damaged Proteins †
Toh, K., Nishio, K., Nakagawa, R., Egoshi, S., Abo, M., Perron, A., Sato, S., Okumura, N., Koizumi, N., Dodo, K., Sodeoka, M., Uesugi, M. †
J. Am. Chem. Soc. 144(44), 20171–20176 (2022)
https://doi.org/10.1021/jacs.2c07180
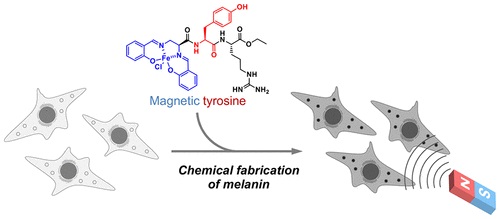
Magnetic Control of Cells by Chemical Fabrication of Melanin †
Nishio, K., Toh, K., Perron, A., Goto, M., Abo, M., Shimakawa, Y., Uesugi, M. †
J. Am. Chem. Soc. 144(37), 16720–16725 (2022)
https://doi.org/10.1021/jacs.2c06555
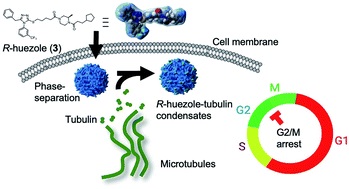
Discovery of a phase-separating small molecule that selectively sequesters tubulin in cells †
Ado, G., Noda, N., Vu, H., Perron, A., Mahapatra, A., Arista, K., Yoshimura, H., Packwood, D., Ishidate, F., Sato, S., Ozawa, T., Uesugi, M. †
Chemical Science 13, 5760-5766 (2022)
https://doi.org/10.1039/D1SC07151C
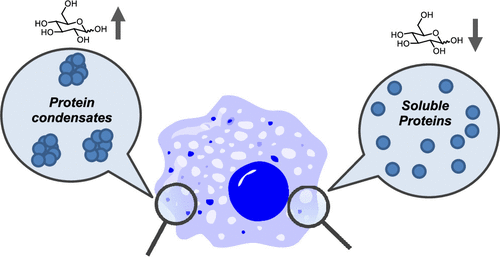
Glucose as a Protein-Condensing Cellular Solute †
Noda, N., Jung, Y., Ado, G., Mizuhata, Y., Higuchi, M., Ogawa, T., Ishidate, F., Sato, S., Kurata, H., Tokitoh, N., Uesugi, M. †
ACS Chem. Biol. 17(3), 567–575 (2022)
https://doi.org/10.1021/acschembio.1c00849
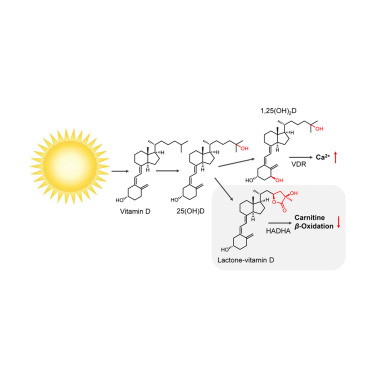
Controlled lipid β-oxidation and carnitine biosynthesis by a vitamin D metabolite. †
Mendoza, A., Takemoto, Y., Cruzado, K., Masoud, S., Nagata, A., Tantipanjaporn, A., Okuda, S., Kawagoe, F., Sakamoto, R., Odagi, M., Mototani, S., Togashi, M., Kawatani, M., Aono, H., Osada, H., Nakagawa, H., Higashi, T., Kittaka, A., Nagasawa, K., Uesugi, M. †
Cell Chemical Biology 29(4), 660-669 (2022)
https://doi.org/10.1016/j.chembiol.2021.08.008
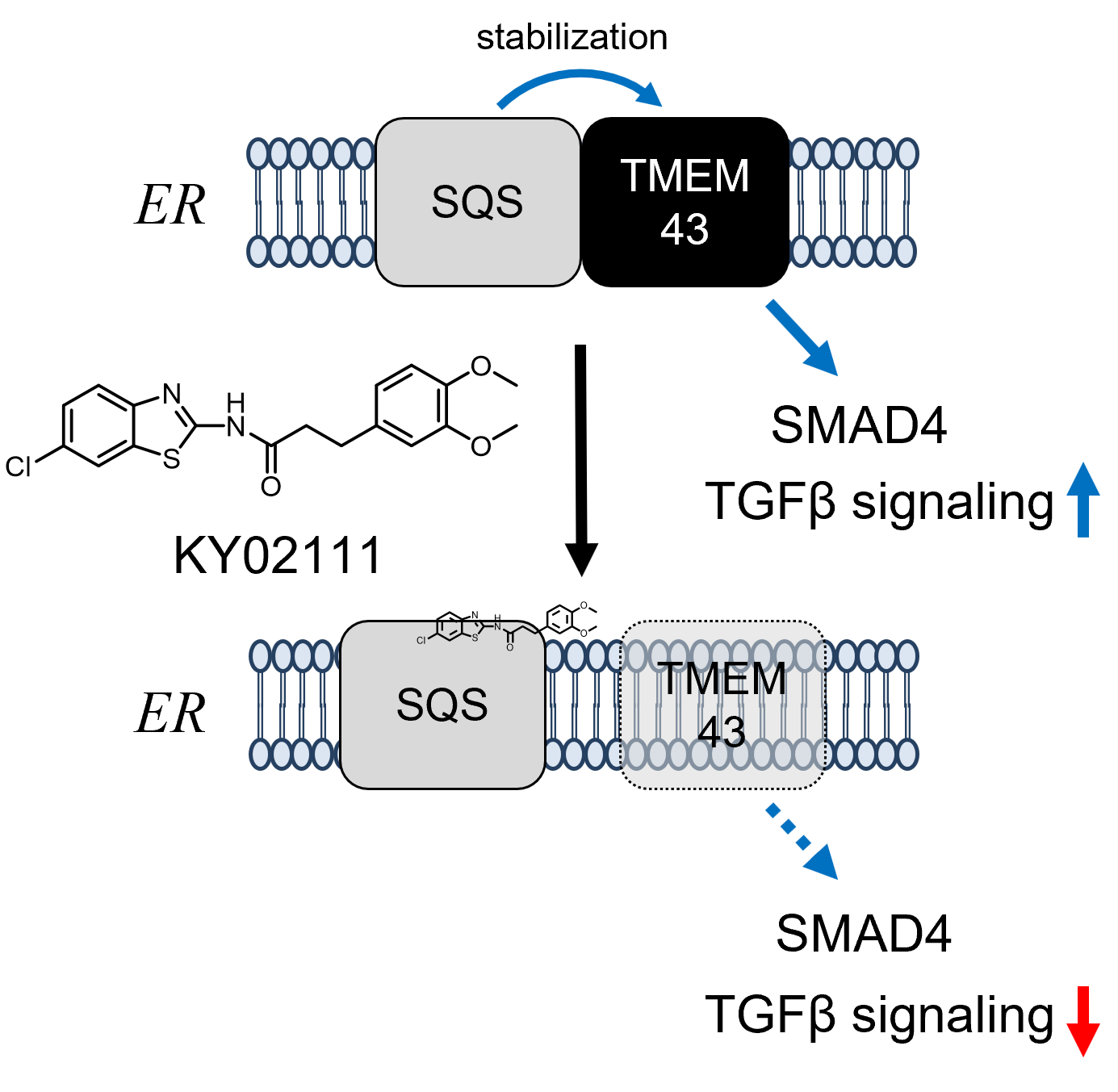
Chemical Genetics Reveals a Role of Squalene Synthase in TGFβ Signaling and Cardiomyogenesis †
Takemoto, Y., Kadota, S., Minami, I., Otsuka, S., Okuda, S., Abo, M., Punzalan, L., Shen, Y., Shiba, Y., Uesugi, M. †
Angew. Chem. Int. Ed. 60(40), 21824-21831 (2021)
https://doi.org/10.1002/anie.202100523
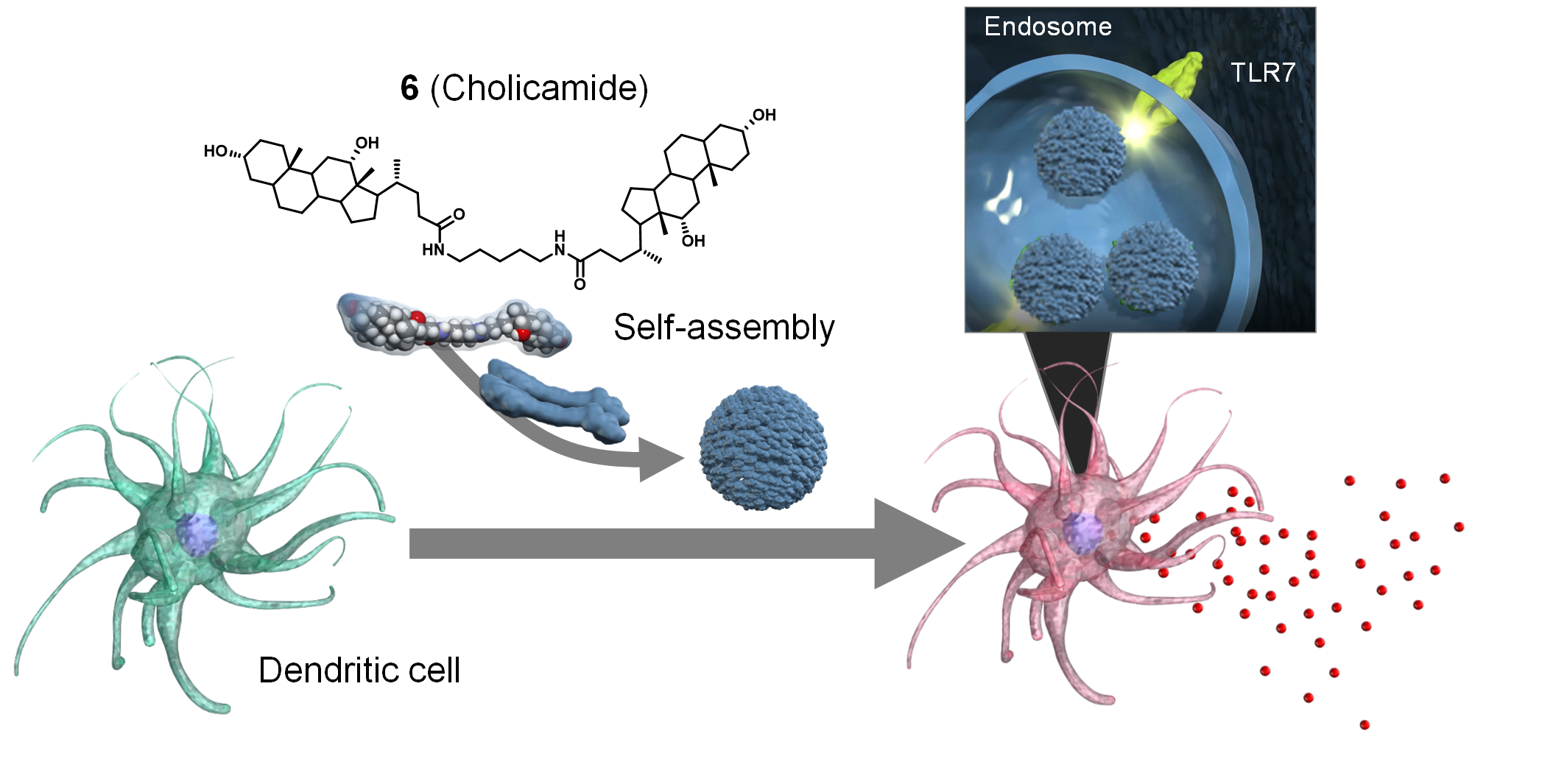
Discovery of Self‐Assembling Small Molecules as Vaccine Adjuvants. †
Jin, S., Vu, H., Hioki, K., Noda, N., Yoshida, H., Shimane, T., Ishizuka, S., Takashima, I., Mizuhata, Y., Pe, K., Ogawa, T., Nishimura, N., Packwood, D., Tokitoh, N., Kurata, H., Yamasaki, S., Ishii, K., Uesugi, M. †
Angew. Chem. Int. Ed. 60(2), 961-969 (2021)
https://doi.org/10.1002/anie.202011604
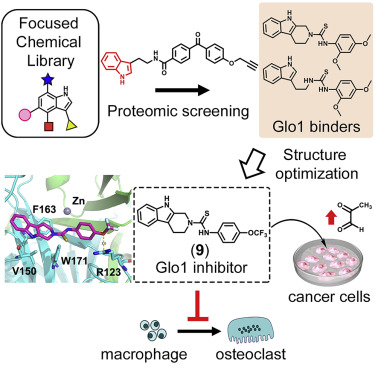
Chemoproteomic Profiling of a Pharmacophore-Focused Chemical Library. †
Punzalan, L., Jiang, L., Mao, D., Mahapatra, A., Sato, S., Takemoto, Y., Tsujimura, M., Kusamori, K., Nishikawa, M., Zhou, L., Uesugi, M. †
Cell Chemical Biology 27(6), 708–718 (2020)
https://doi.org/10.1016/j.chembiol.2020.04.007
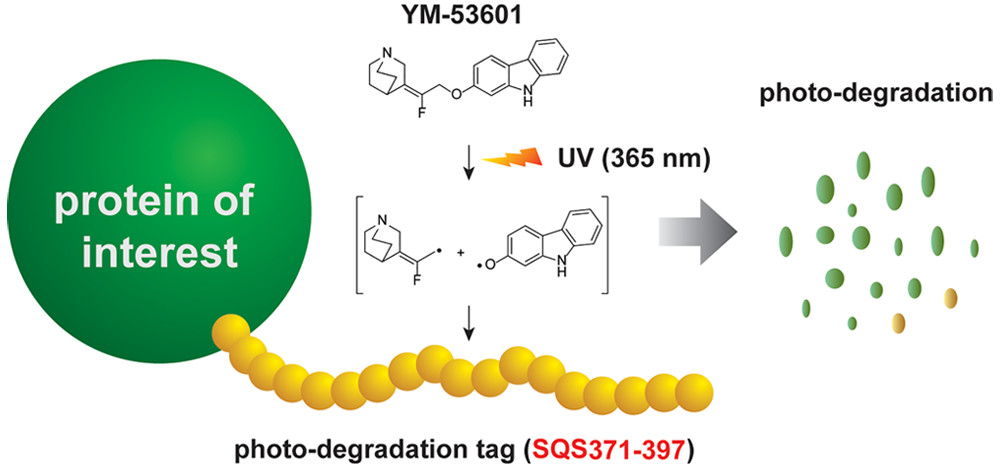
Discovery of a Small-Molecule-Dependent Photolytic Peptide. †
Takemoto, Y., Mao, Di., Punzalan, L., Götze, S., Sato, S., Uesugi, M. †
J. Am. Chem. Soc. 142(3), 1142-1146 (2020)
https://doi.org/10.1021/jacs.9b09178
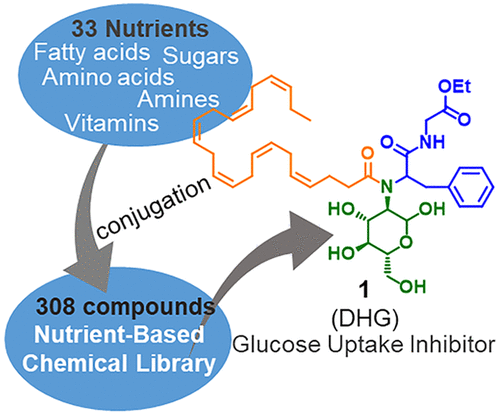
Nutrient-Based Chemical Library as a Source of Energy Metabolism Modulators. †
Furuta,T., Mizukami, Y., Asano, L., Kotake, K., Ziegler, S., Yoshida, H., Watanabe, M., Sato, S., Waldmann, H., Nishikawa, M., Uesugi, M. †
ACS Chem. Biol. 14(9), 1860-1865 (2019)
https://doi.org/10.1021/acschembio.9b00444
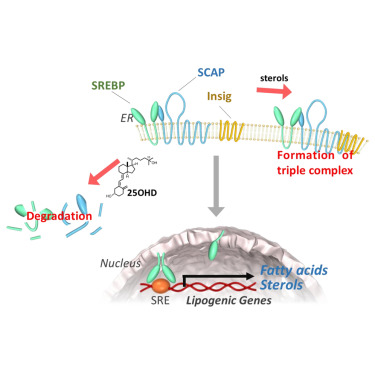
Vitamin D metabolite, 25-Hydroxyvitamin D, regulates lipid metabolism by inducing degradation of SREBP/SCAP. †
Asano, L., Watanabe, M., Ryoden, Y., Usuda, K., Yamaguchi, T., Khambu, B., Takashima, M., Sato, S., Sakai, J., Nagasawa, K., Uesugi, M. †
Cell Chem Biol. 24, 207-217 (2017)
https://doi.org/10.1016/j.chembiol.2016.12.017
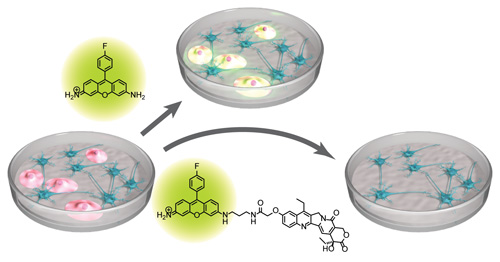
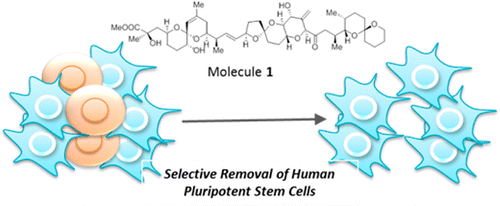
A synthetic hybrid molecule for the selective removal of human pluripotent stem cells from cell mixtures. †
Mao, D., Ando, S., Sato, S., Qin, Y., Hirata, N., Katsuda, Y., Kawase, E., Kuo, T.F., Minami, I., Shiba, Y., Ueda, K., Nakatsuji, N., Uesugi, M. †
Angew. Chem. Int. Ed. 56(7), 1765-1770 (2017)
https://doi.org/10.1002/anie.201610284
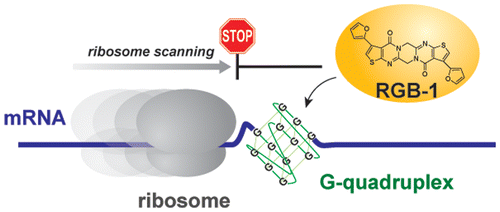
A small molecule that represses translation of G-quadruplex-containing mRNA. †
Katsuda, Y., Sato, S., Asano, L., Morimura, Y., Furuta, T., Sugiyama, H., Hagihara, M., Uesugi, M. †
J. Am. Chem. Soc. 138, 9037-9040 (2016)
https://doi.org/10.1021/jacs.6b04506
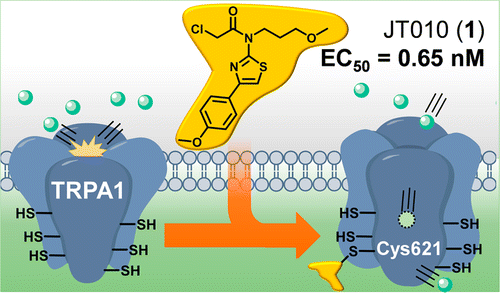
A potent and site-selective agonist of TRPA1. †
Takaya, J., Mio, K., Shiraishi, T., Kurokawa, T., Otsuka, S., Mori, Y., Uesugi, M. †
J. Am. Chem. Soc. 137, 15859−15864 (2015)
https://doi.org/10.1021/jacs.5b10162
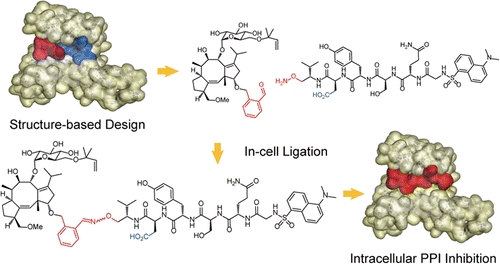
Intracellular generation of a diterpene-peptide conjugate that inhibits 14-3-3-mediated interactions. †
Parvatkar, P., Kato, N., Uesugi, M., Sato, S., Ohkanda, J. †
J. Am. Chem. Soc. 137, 15624-15627 (2015)
https://doi.org/10.1021/jacs.5b09817
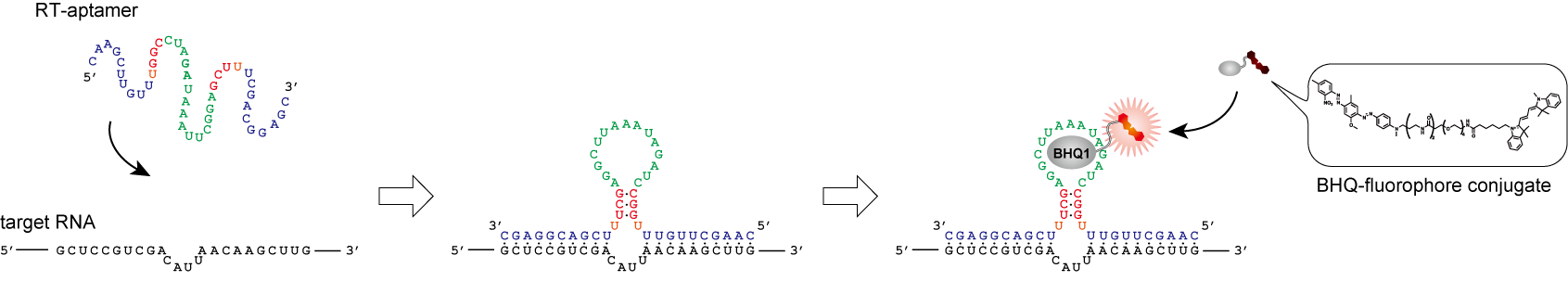
Live-cell imaging of endogenous mRNAs with a small molecule. †
Sato, S., Watanabe, M., Katsuda, Y., Murata, A., Wang, D. O., Uesugi M. †
Angew. Chem. Int. Ed. 54(6), 1855-1858 (2015)
https://doi.org/10.1002/anie.201410339
Synthetic molecules that protect cells from anoikis and their use in cell transplantation. †
Frisco-Cabanos, H.L., Watanabe, M., Okumura, N., Kusamori, K., Takemoto, N., Takaya, J., Sato, S., Yamazoe, S., Takakura, Y., Kinoshita, S., Nishikawa, M., Koizumi, N., Uesugi, M. †
Angew. Chem. Int. Ed. 53(42), 11208-11213 (2014)
https://doi.org/10.1002/anie.201405829
Selective elimination of human pluripotent stem cells by a marine natural product derivative. †
Kuo, T.F., Mao, D., Hirata, N., Khambu, B., Kimura, Y., Kawase, E., Shimogawa, H., Ojika, M., Nakatsuji, N., Ueda, K., Uesugi, M. †
J. Am. Chem. Soc. 136 (28), 9798-9801 (2014)
https://doi.org/10.1021/ja501795c
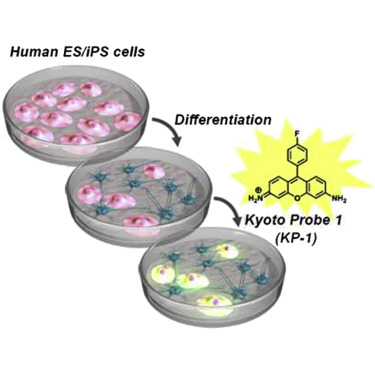
A chemical probe that labels human pluripotent stem cells. †
Hirata, N., Nakagawa, M., Fujibayashi, Y., Yamauchi, K., Murata, A., Minami, I., Tomioka, M., Kondo, T., Kuo, T.F., Endo, H., Inoue, H., Sato, S., Ando, S., Kawazoe, Y., Aiba, K., Nagata, K., Kawase, E., Chang, Y.T., Suemori, H., Eto, K., Nakauchi, H., Yamanaka, S., Nakatsuji, N., Ueda, K., Uesugi, M. †
Cell Rep 6., 1165–1174 (2014)
https://doi.org/10.1016/j.celrep.2014.02.006
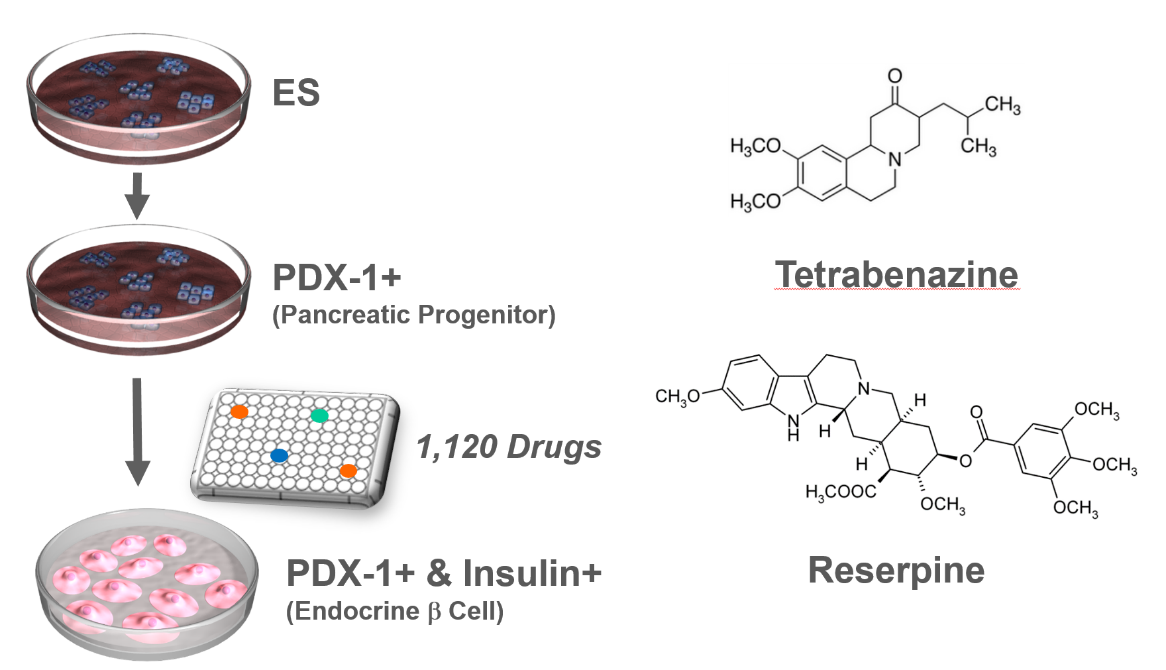
VMAT2 identified as a regulator of late-stage beta cell differentiation. †
Sakano, D., Shiraki, N., Kikawa, K., Yamazoe, T., Kataoka, M., Umeda, K., Araki, K., Mao, D., Matsumoto, S., Nakagata, N., Andersson, O., Stainier, D., Endo, F., Kume, K., Uesugi, M., Kume, S. †
Nat. Chem. Biol. 10, 141-148 (2014)
https://doi.org/10.1038/nchembio.1410
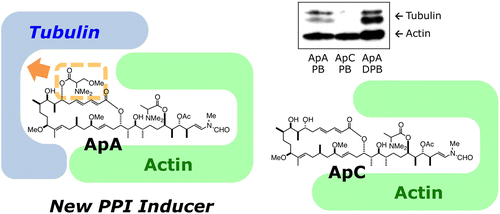
Inhibition of microtubule assembly by a complex of actin and antitumor macrolide aplyronine A. †
Kita, M., Hirayama, Y., Yoneda, K., Yamagishi, K., Chinen, T., Usui, T., Sumiya, E., Uesugi, M., Kigoshi, H. †
J. Am. Chem. Soc. 135 (48), 18089-18095 (2013)
https://doi.org/10.1021/ja406580w
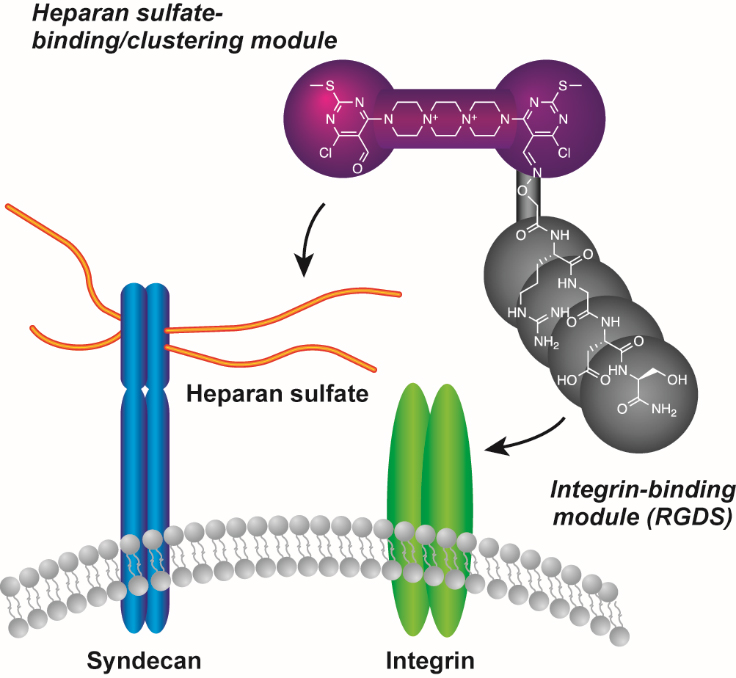
Small molecule-induced clustering of heparan sulfate promotes cell adhesion. †
Takemoto, N., Suehara, T., Frisco, H., Sato, S., Sezaki, T., Kusamori, K., Kawazoe, Y., Park, S., Yamazoe, S., Mizuhata, Y., Inoue, R., Miller, G., Hansen, S., Jayson, G., Gardiner, J., Kanaya, T., Tokitoh, N., Ueda, K., Takakura, Y., Kioka, N., Nishikawa, M., Uesugi, M. †
J. Am. Chem. Soc. 135 (30), 11032-11039 (2013)
https://doi.org/10.1021/ja4018682
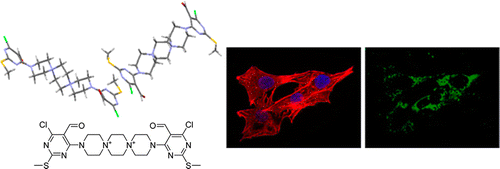
A small molecule that promotes cardiac differentiation of human pluripotent stem cells under defined cytokine- and xeno-free conditions. †
Minami, I., Yamada, K., Otsuji, T.G., Yamamoto, T., Shen, Y., Otsuka, S., Kadota, S., Morone, N., Barve, M., Asai, Y., Tenkova-Heuser, T., Heuser, J. E., Uesugi, M., Aiba, K., Nakatsuji, N. †
Cell Rep. (5), 1448-1460 (2012)
https://doi.org/10.1016/j.celrep.2012.09.015
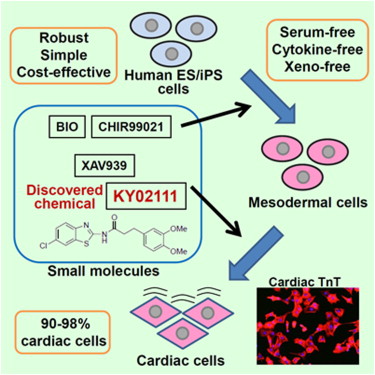
Synthesis and evaluation of diarylthiazole derivatives that inhibit activation of sterol regulatory element-binding proteins. †
Kamisuki, S., Shirakawa T., Kugimiya, A., Abu-Elheiga, L., Choo, H.-Y., Yamada, K., Shimogawa, H., Wakil, S. J., Uesugi, M. †
J. Med. Chem. 54(13), 4923-4927 (2011)
https://doi.org/10.1021/jm200304y
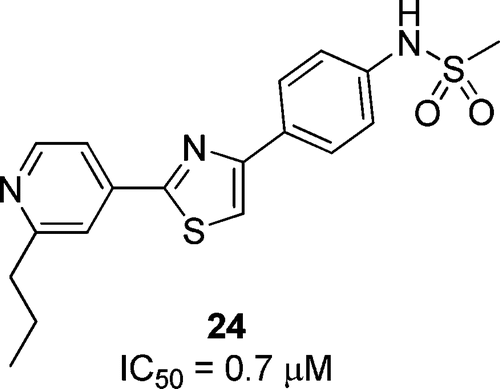
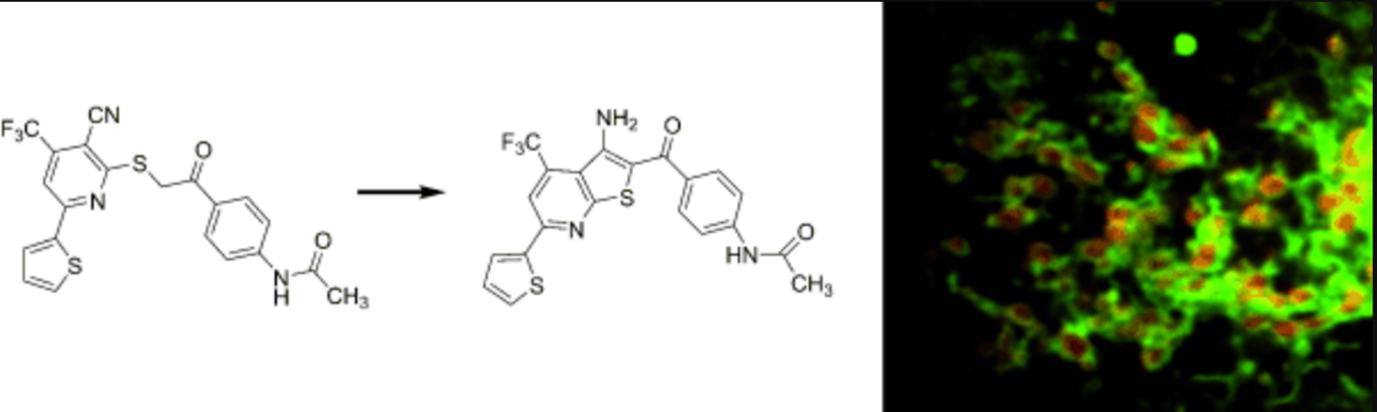
A mitochondrial surface-specific fluorescent probe activated by bioconversion. †
Kawazoe, Y., Shimogawa, H., Sato, A., Uesugi, M. †
Angew. Chem. Int. Ed. 50(24), 5478-5481 (2011)
https://doi.org/10.1002/anie.201100935
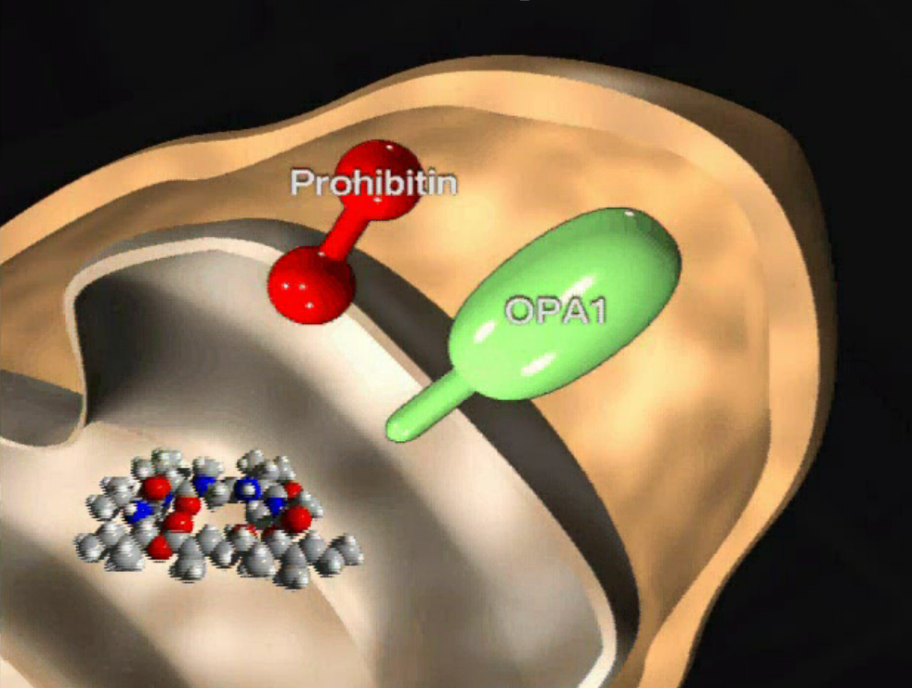
Marine natural product aurilide activates the OPA1-mediated apoptosis by binding to prohibitin. †
Sato, S. Murata, A., Orihara, T., Shirakawa, T., Suenaga, K., Kigoshi, H., Uesugi, M. †
Chem. Biol. 18 (1), 131-139 (2011)
https://doi.org/10.1016/j.chembiol.2010.10.017
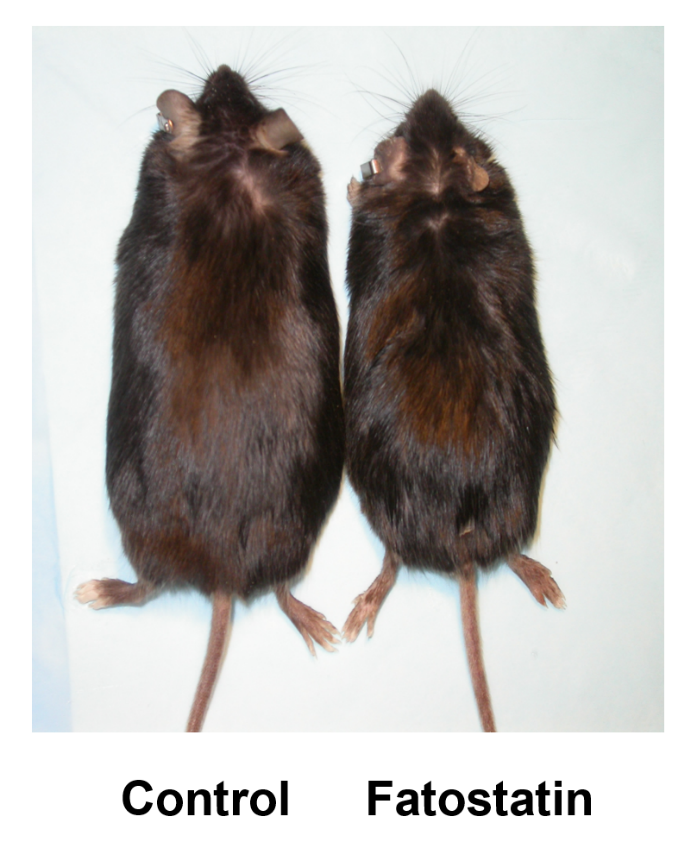
A small molecule that blocks fat synthesis by inhibiting the activation of SREBP. †
Kamisuki, S., Mao, Q., Abu-Eliheiga, L., Gu, Z., Kugimiya, A., Kwon, Y., Shinohara, T., Kawazoe, Y., Sato, S. Asakura, K., Choo, H., Sakai, J., Wakil, SJ., Uesugi, M. †
Chem. Biol. 16 (8), 882-892 (2009)
https://doi.org/10.1016/j.chembiol.2009.07.007
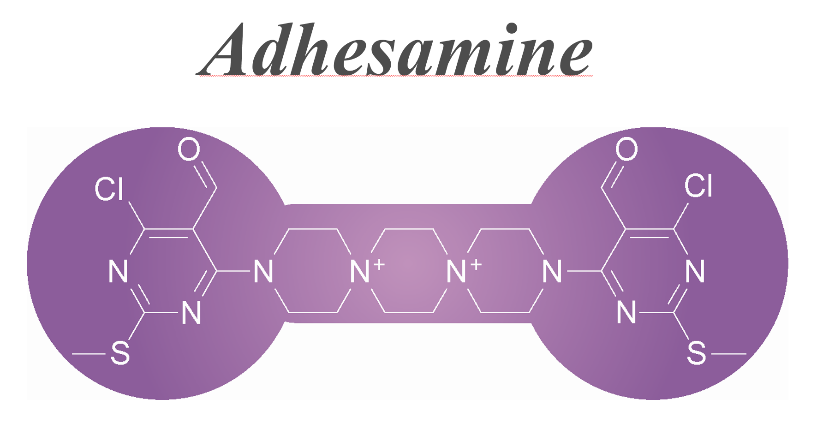
A dumbbell-shaped small molecule that promotes cell adhesion and growth. †
Yamazoe, S., Shimogawa, H., Sato, S., Esko, J. D., Uesugi, M. †
Chem. Biol. 16 (7), 773-782 (2009)
https://doi.org/10.1016/j.chembiol.2009.06.008
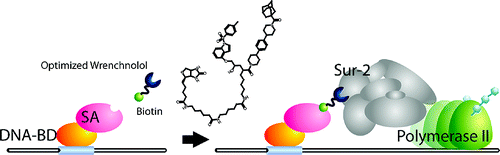
Wrenchnolol derivative optimized for gene activation in cells. †
Jung, D., Shimogawa, H., Kwon, Y., Mao, Q., Sato, S., Kamisuki, S., Kigoshi, H., Uesugi, M. †
J. Am. Chem. Soc. 131(13), 4774-4782 (2009)
https://doi.org/10.1021/ja900669k
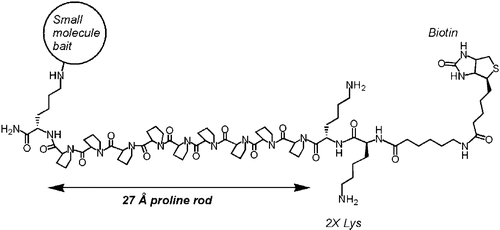
Polyproline-rod approach to isolating protein targets of bioactive small molecules: isolation of a new target of indomethacin. †
Sato, S., Kwon, Y., Kamisuki, S., Srivastava, N., Mao, Q., Kawazoe, Y., Uesugi, M. †
J. Am. Chem. Soc. 129(4), 873-880 (2007)
https://doi.org/10.1021/ja0655643
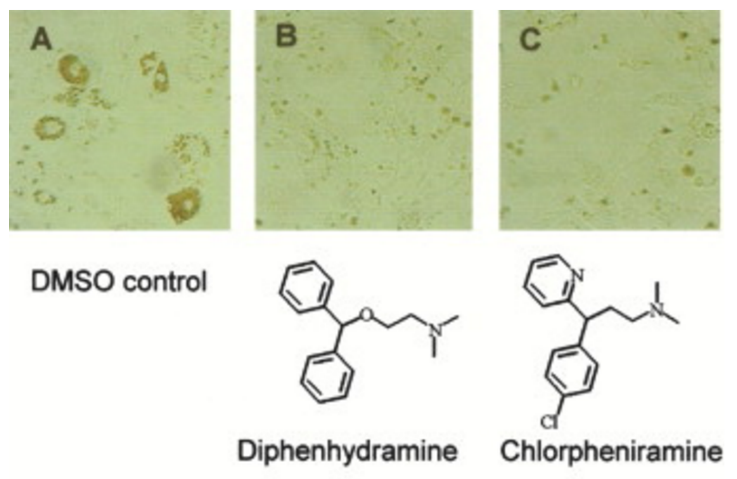
Chemical genetic identification of histamine receptor H1 as a stimulator of insulin-induced adipogenesis. †
Kawazoe, Y., Tanaka, H., Uesugi, M. †
Chem. Biol. 11, 907-913 (2004)
https://doi.org/10.1016/j.chembiol.2004.04.017
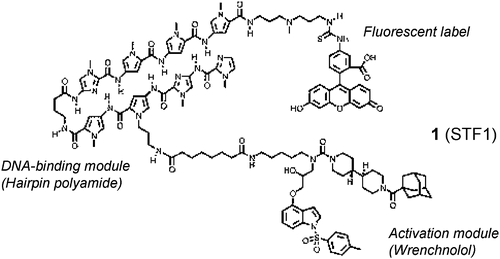
Small molecule transcription factor mimic. †
Kwon,Y., Arndt, H., Mao, Q., Choi, Y., Kawazoe, Y., Dervan, P. B., Uesugi, M. †
J. Am. Chem. Soc. 126, 15940-15941 (2004)
https://doi.org/10.1021/ja0445140
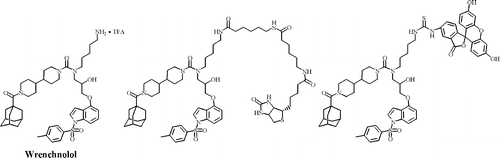
A wrench-shaped synthetic molecule that modulates a transcription factor-coactivator interaction. †
Shimogawa, H., Kwon, Y., Mao, Q., Kawazoe, Y., Choi, Y., Asada, S., Kigoshi, H., & Uesugi, M. †
J. Am. Chem. Soc. 126, 3461-3471 (2004)
https://doi.org/10.1021/ja038855+
A gene-expression inhibitor that targets an alpha-helix-mediated protein interaction. †
Asada, S., Choi, Y., & Uesugi, M. †
J. Am. Chem. Soc. 125, 4992-4993 (2003)
https://doi.org/10.1021/ja0292703
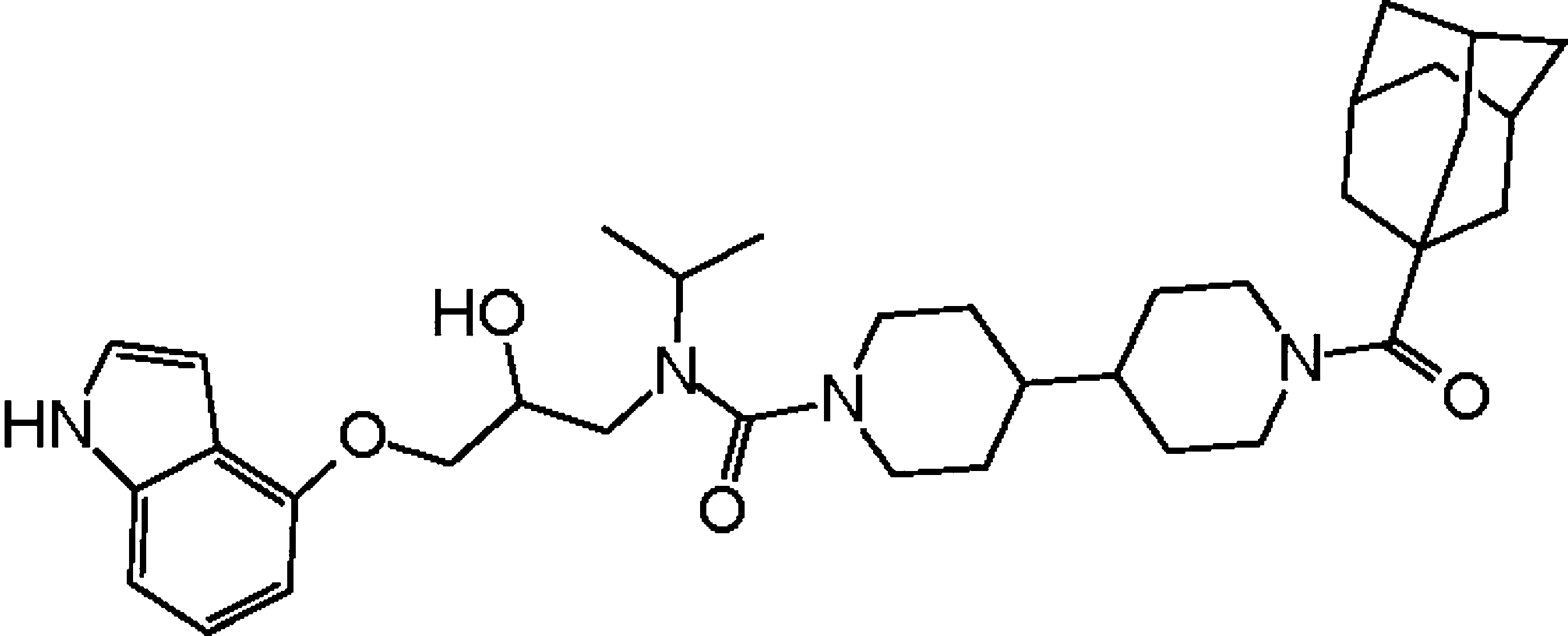
External control of Her2 expression and cancer cell growth by targeting a Ras-linked coactivator. †
Asada, S., Choi, Y., Yamada, M., Wang, S., Hung, M., Qin, J., & Uesugi, M. †
Proc. Natl. Acad. Sci. USA 99, 12747-12752 (2002)
https://doi.org/10.1073/pnas.202162199
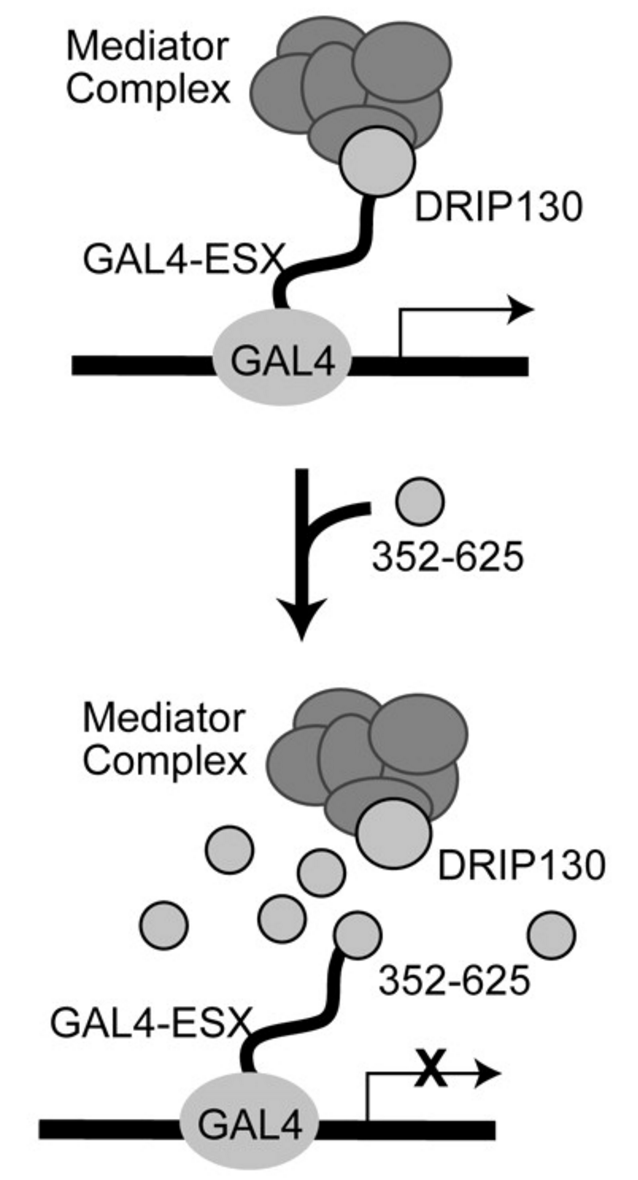
Divergent hTAF(II)31-binding motifs hidden in activation domains. †
Choi, Y., Asada, S., & Uesugi, M. †
J. Biol. Chem. 275, 15912-15916 (2000)
https://doi.org/10.1074/jbc.275.21.15912

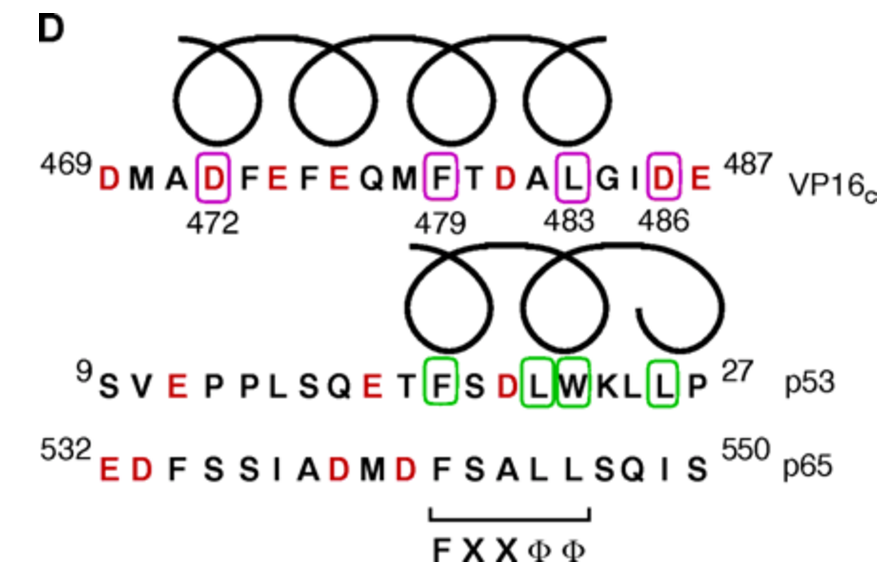
The alpha helical FXXphiphi motif in p53: TAF interaction and discrimination by MDM2. †
Uesugi, M. & Verdine, G. L. †
Proc. Natl. Acad. Sci. USA 96, 14801-14806 (1999)
https://doi.org/10.1073/pnas.96.26.14801
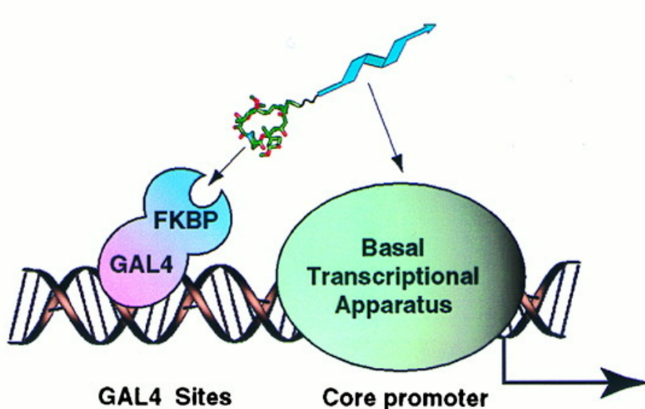
A non-natural transcriptional coactivator. †
Nyanguile, O., Uesugi, M., Austin, D. J., & Verdine, G. L. †
Proc. Natl. Acad. Sci. USA 94, 13402-13406 (1997)
https://doi.org/10.1073/pnas.94.25.13402
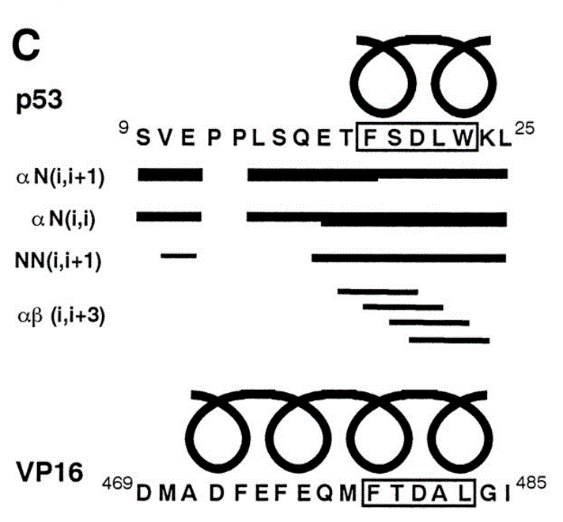
Induced alpha helix in the VP16 activation domain upon binding to a human TAF. †
Uesugi, M., Nyanguile, O., Lu, H., Levine, A. J., & Verdine, G. L. †
Science 277, 1310-1313 (1997)
https://doi.org/10.1126/science.277.5330.1310